Synthetic Control Method
This notebook presents the in store navigation case and the core analysis workflow.
Data
Ground-truth ATTE is 2.750000
| unit_id | calendar_time | treated_time | y | y_cf | tau_realized_true | mu_cf | mu_treated | tau_mean_true | |
|---|---|---|---|---|---|---|---|---|---|
| 0 | donor_1 | 2000-01 | 0 | 4.0 | 4.0 | 0.0 | 6.150671 | 6.150671 | 0.0 |
| 1 | donor_1 | 2000-02 | 0 | 9.0 | 9.0 | 0.0 | 5.599403 | 5.599403 | 0.0 |
| 2 | donor_1 | 2000-03 | 0 | 4.0 | 4.0 | 0.0 | 4.978119 | 4.978119 | 0.0 |
| 3 | donor_1 | 2000-04 | 0 | 6.0 | 6.0 | 0.0 | 5.777247 | 5.777247 | 0.0 |
| 4 | donor_1 | 2000-05 | 0 | 8.0 | 8.0 | 0.0 | 5.685711 | 5.685711 | 0.0 |
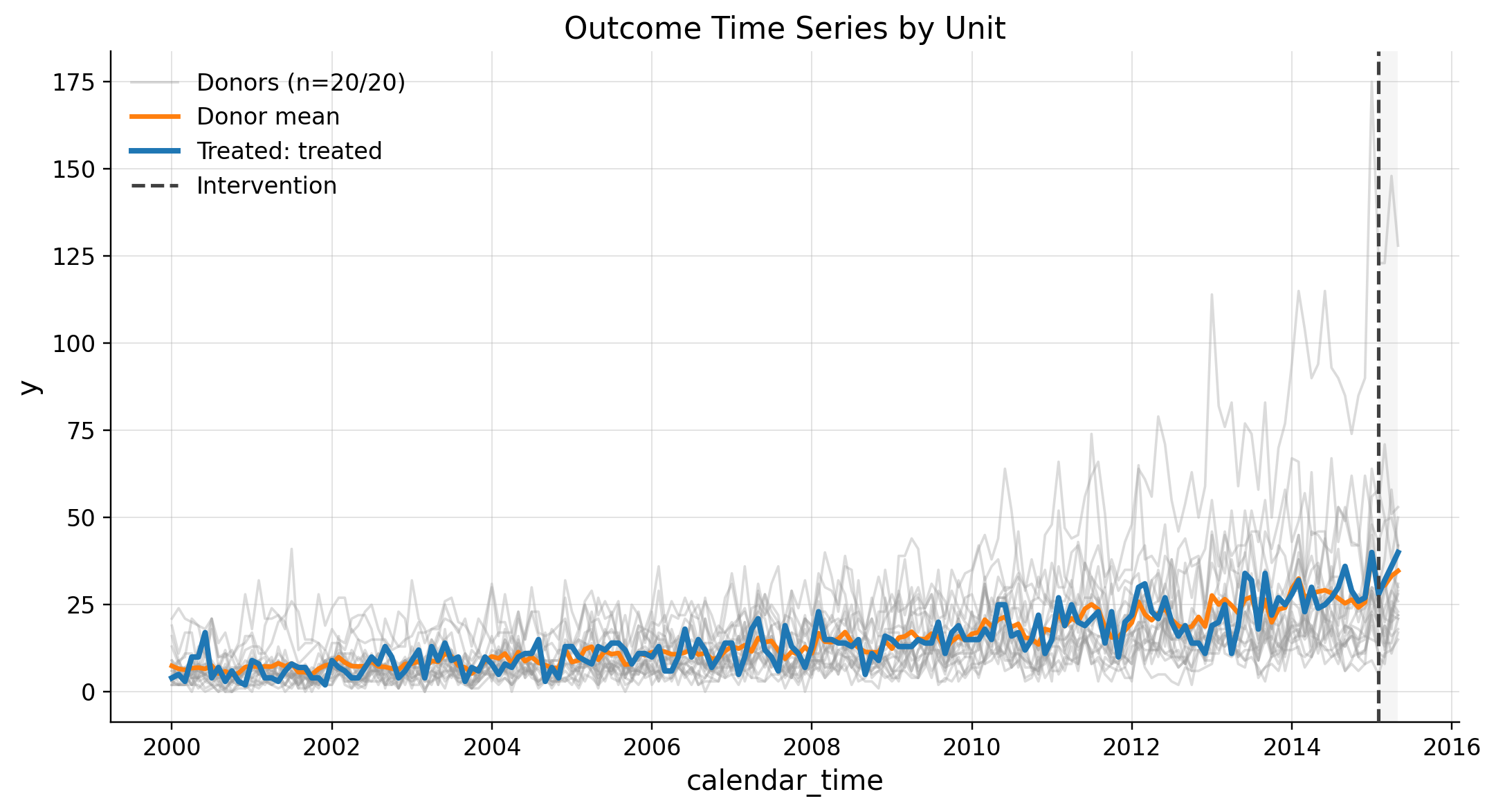
EDA
PanelDataSCM(df=(3885, 4), y='y', unit_col='unit_id', time_col='calendar_time', treated_time='treated_time', time_freq='M', treated_unit='treated', treatment_start=Period('2015-02', 'M'), last_post_period=Period('2015-05', 'M'), n_pre_periods=181, n_post_periods=4, donor_units=['donor_1', 'donor_10', 'donor_11', 'donor_12', 'donor_13', 'donor_14', 'donor_15', 'donor_16', 'donor_17', 'donor_18', 'donor_19', 'donor_2', 'donor_20', 'donor_3', 'donor_4', 'donor_5', 'donor_6', 'donor_7', 'donor_8', 'donor_9'])
| donor | pre_mean | pre_std | pre_slope | corr_with_treated_pre | rmse_to_treated_pre | rmse_to_treated_pre_standardized | mean_diff_pre | slope_diff_pre | max_abs_gap_pre | is_never_treated | n_missing_pre | corr_rank | std_rmse_rank | slope_rank | composite_similarity_score | rank_by_similarity | notes | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | donor_4 | 9.762431 | 7.179154 | 0.115553 | 0.775096 | 6.670625 | 0.842433 | -4.287293 | -0.007929 | 20.0 | True | 0 | 4 | 5 | 1 | 0.883333 | 1 | ok |
| 1 | donor_10 | 13.563536 | 7.602892 | 0.114674 | 0.755004 | 5.462074 | 0.689805 | -0.486188 | -0.008807 | 17.0 | True | 0 | 8 | 1 | 2 | 0.866667 | 2 | ok |
| 2 | donor_18 | 16.220994 | 9.298357 | 0.140931 | 0.774548 | 6.310125 | 0.796906 | 2.171271 | 0.017449 | 20.0 | True | 0 | 5 | 3 | 3 | 0.866667 | 3 | ok |
| 3 | donor_9 | 13.701657 | 6.733231 | 0.100152 | 0.731460 | 5.491832 | 0.693564 | -0.348066 | -0.023330 | 19.0 | True | 0 | 10 | 2 | 4 | 0.783333 | 4 | ok |
| 4 | donor_1 | 10.646409 | 5.490006 | 0.079764 | 0.740206 | 6.329794 | 0.799390 | -3.403315 | -0.043717 | 20.0 | True | 0 | 9 | 4 | 6 | 0.733333 | 5 | ok |
| 5 | donor_17 | 17.458564 | 10.886897 | 0.178779 | 0.770667 | 7.744183 | 0.978013 | 3.408840 | 0.055297 | 27.0 | True | 0 | 6 | 9 | 9 | 0.650000 | 6 | ok |
| 6 | donor_19 | 10.453039 | 5.153367 | 0.073788 | 0.691311 | 6.765610 | 0.854429 | -3.596685 | -0.049693 | 23.0 | True | 0 | 12 | 6 | 7 | 0.633333 | 7 | ok |
| 7 | donor_13 | 8.718232 | 5.794182 | 0.080950 | 0.681039 | 7.886978 | 0.996047 | -5.331492 | -0.042531 | 30.0 | True | 0 | 13 | 10 | 5 | 0.583333 | 8 | ok |
| 8 | donor_8 | 9.198895 | 5.152288 | 0.070184 | 0.669963 | 7.623024 | 0.962712 | -4.850829 | -0.053298 | 23.0 | True | 0 | 14 | 8 | 8 | 0.550000 | 9 | ok |
| 9 | donor_5 | 10.950276 | 5.267723 | 0.065347 | 0.609616 | 7.014192 | 0.885823 | -3.099448 | -0.058134 | 22.0 | True | 0 | 16 | 7 | 10 | 0.500000 | 10 | ok |
| 10 | donor_7 | 7.204420 | 4.610483 | 0.064872 | 0.696173 | 8.943345 | 1.129455 | -6.845304 | -0.058610 | 22.0 | True | 0 | 11 | 12 | 11 | 0.483333 | 11 | high_std_rmse |
| 11 | donor_11 | 21.038674 | 12.944576 | 0.224528 | 0.828828 | 10.449828 | 1.319709 | 6.988950 | 0.101046 | 35.0 | True | 0 | 2 | 16 | 17 | 0.466667 | 12 | high_std_rmse |
| 12 | donor_14 | 15.685083 | 15.073494 | 0.250056 | 0.832498 | 9.687949 | 1.223491 | 1.635359 | 0.126574 | 34.0 | True | 0 | 1 | 15 | 19 | 0.466667 | 13 | high_std_rmse |
| 13 | donor_6 | 8.690608 | 4.149602 | 0.049649 | 0.564090 | 8.459850 | 1.068395 | -5.359116 | -0.073833 | 27.0 | True | 0 | 17 | 11 | 13 | 0.366667 | 14 | high_std_rmse |
| 14 | donor_12 | 7.569061 | 3.764025 | 0.046441 | 0.618555 | 9.055080 | 1.143566 | -6.480663 | -0.077040 | 31.0 | True | 0 | 15 | 13 | 14 | 0.350000 | 15 | high_std_rmse |
| 15 | donor_15 | 27.751381 | 13.661215 | 0.212821 | 0.764745 | 16.480694 | 2.081347 | 13.701657 | 0.089339 | 53.0 | True | 0 | 7 | 19 | 16 | 0.350000 | 16 | high_std_rmse |
| 16 | donor_2 | 31.861878 | 29.367320 | 0.482832 | 0.825508 | 29.299935 | 3.700290 | 17.812155 | 0.359351 | 135.0 | True | 0 | 3 | 20 | 20 | 0.333333 | 17 | high_std_rmse |
| 17 | donor_20 | 19.972376 | 6.508330 | 0.056204 | 0.480955 | 9.516488 | 1.201837 | 5.922652 | -0.067278 | 23.0 | True | 0 | 19 | 14 | 12 | 0.300000 | 18 | high_std_rmse |
| 18 | donor_16 | 5.662983 | 3.652322 | 0.040390 | 0.561087 | 10.673455 | 1.347951 | -8.386740 | -0.083091 | 31.0 | True | 0 | 18 | 17 | 15 | 0.216667 | 19 | high_std_rmse |
| 19 | donor_3 | 19.900552 | 5.562408 | -0.000138 | 0.105856 | 10.887927 | 1.375037 | 5.850829 | -0.123619 | 33.0 | True | 0 | 20 | 18 | 18 | 0.116667 | 20 | high_std_rmse |
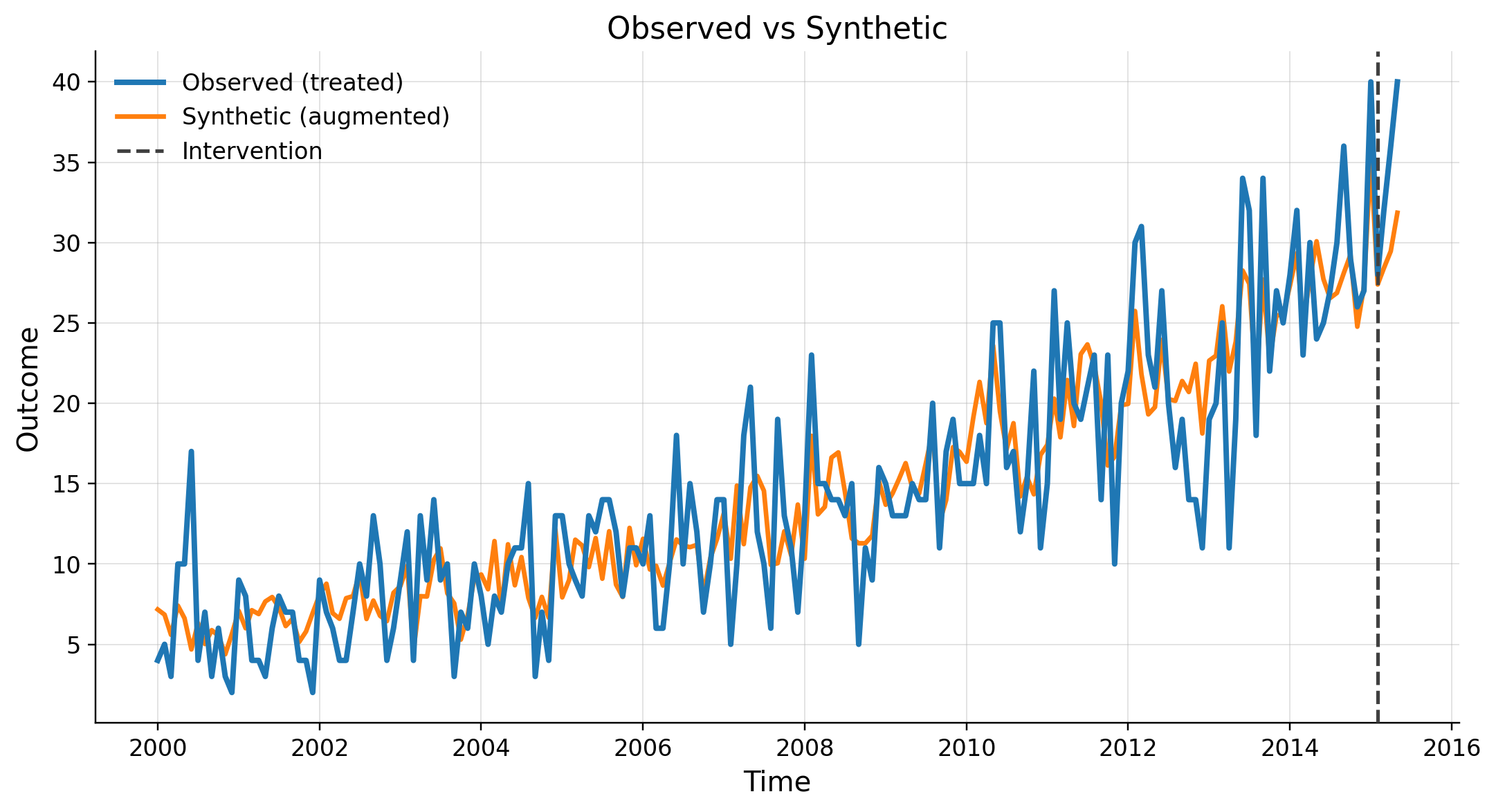
Inference
| value | |
|---|---|
| field | |
| estimand | average_post_effect |
| model | AugmentedSyntheticControl |
| inference | average_att_ttest |
| value | 4.9727 (ci_abs: 0.2503, 9.6951) |
| value_relative | 16.9843 (ci_rel: 0.8551, 33.1135) |
| alpha | 0.0500 |
| p_value | 0.0411 |
| is_significant | True |
| post_outcome_d_mean | 34.0000 |
| pointwise_post_period_average | 4.7216 |
| effect_by_time | [{'period': 2015-02, 'estimate': 0.60800479979... |
| cumulative_effect | 18.8864 |
| n_significant_periods | 0 |
| n_donors | 20 |
| n_pre_periods | 181 |
| n_post_periods | 4 |
| time | 2026-03-09 |
Refutation
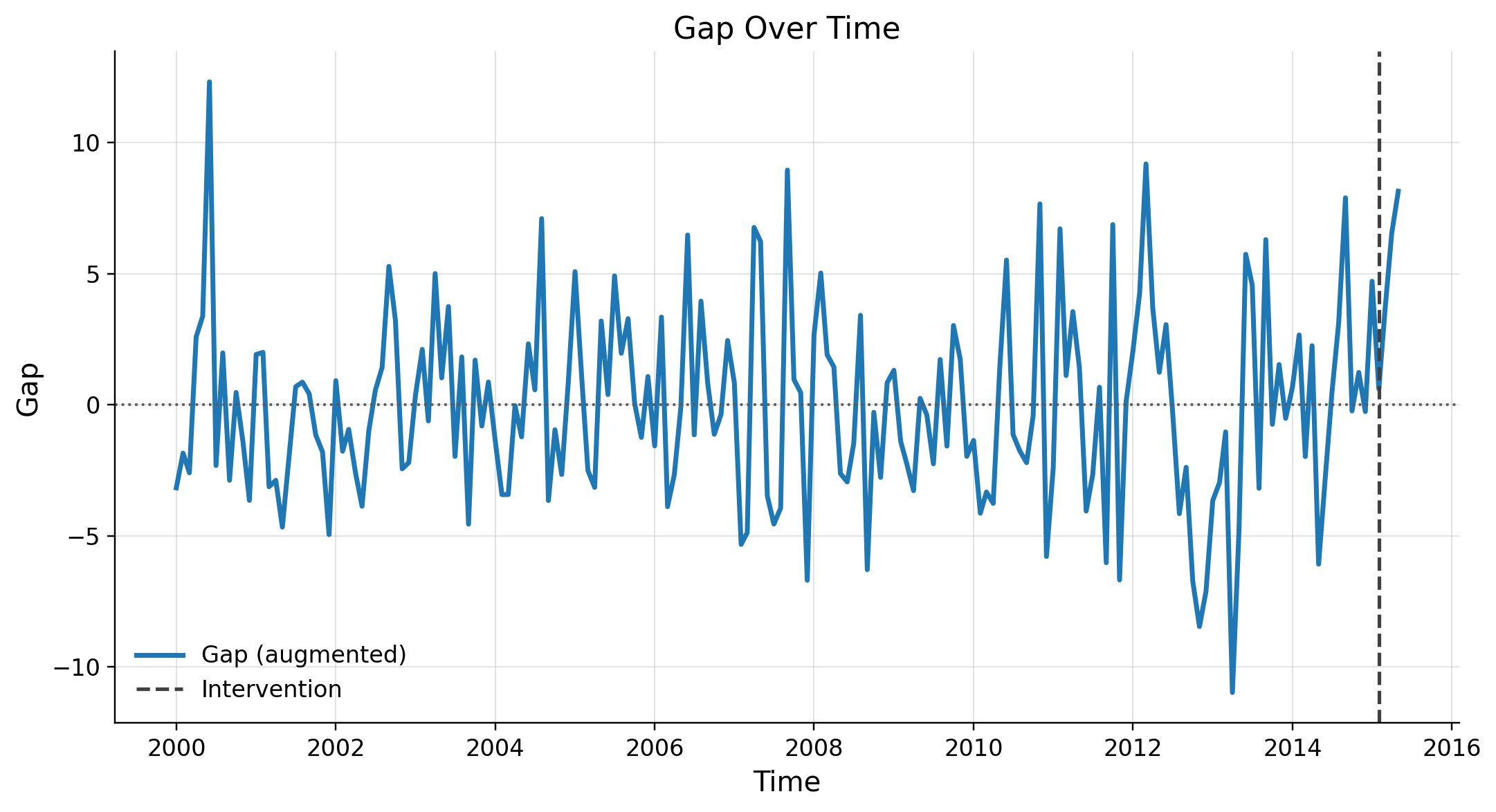
Placebo
| placebo_treatment_start | n_pre_before_placebo | n_post_after_placebo | average_att_placebo | ci_lower | ci_upper | p_value | rejects_zero | pre_fit_metric | |
|---|---|---|---|---|---|---|---|---|---|
| 0 | 2000-03 | 2 | 4 | 5.462732 | -14.408865 | 25.334328 | 0.177510 | False | 3.957005e-09 |
| 1 | 2000-04 | 3 | 4 | 5.820208 | 1.568835 | 10.071582 | 0.027632 | True | 1.342351e-09 |
| 2 | 2000-05 | 4 | 4 | 6.443555 | -0.217886 | 13.104995 | 0.053169 | False | 1.285495e-02 |
| 3 | 2000-06 | 5 | 4 | 4.912309 | -1.310838 | 11.135455 | 0.076833 | False | 9.762317e-03 |
| 4 | 2000-07 | 6 | 4 | -3.505020 | -14.728100 | 7.718059 | 0.311187 | False | 6.010438e-02 |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 171 | 2014-06 | 173 | 4 | 3.033284 | -15.201103 | 21.267671 | 0.548432 | False | 3.604817e+00 |
| 172 | 2014-07 | 174 | 4 | 3.855998 | -14.376369 | 22.088365 | 0.458889 | False | 3.596280e+00 |
| 173 | 2014-08 | 175 | 4 | 4.315601 | -13.943856 | 22.575058 | 0.416190 | False | 3.586448e+00 |
| 174 | 2014-09 | 176 | 4 | 3.386534 | -15.538700 | 22.311768 | 0.521847 | False | 3.584910e+00 |
| 175 | 2014-10 | 177 | 4 | 1.954115 | -15.878151 | 19.786382 | 0.683716 | False | 3.645295e+00 |
176 rows × 9 columns
We pretend treatment has happened earlier.
- placebo_treatment_start - where treatment starts
- n_pre_before_placebo - how many periods were fitted before the treatment
- n_post_after_placebo - how many periods were fitted after the treatment
- average_att_placebo - average_post_effect
- rejects_zero - p_value < alpha
So weak robustness when we have many rows have rejects_zero = false and average_att_placebo is small
| unit_id | pre_rmse | post_rmse | post_pre_rmspe_ratio | average_post_gap | max_abs_post_gap | rank_post_pre_rmspe_ratio | is_actual_treated | |
|---|---|---|---|---|---|---|---|---|
| 0 | donor_2 | 9.908261 | 42.582403 | 4.297667 | 35.069571 | 71.993440 | 1 | False |
| 1 | donor_18 | 4.160427 | 13.911060 | 3.343661 | -9.715085 | 26.366275 | 2 | False |
| 2 | donor_11 | 5.227018 | 17.008344 | 3.253929 | -7.814759 | 22.368382 | 3 | False |
| 3 | donor_15 | 7.267022 | 19.260087 | 2.650341 | -11.741052 | 36.900028 | 4 | False |
| 4 | donor_1 | 3.161121 | 7.777379 | 2.460323 | -2.925918 | 12.693281 | 5 | False |
| 5 | donor_14 | 5.041892 | 12.241623 | 2.427982 | 0.209326 | 16.893802 | 6 | False |
| 6 | donor_7 | 2.922371 | 6.708425 | 2.295542 | -0.124026 | 9.677678 | 7 | False |
| 7 | donor_12 | 2.638915 | 5.973629 | 2.263668 | 3.032937 | 10.284256 | 8 | False |
| 8 | donor_4 | 3.281984 | 6.259385 | 1.907195 | 1.824818 | 8.201755 | 9 | False |
| 9 | donor_8 | 3.234938 | 5.850849 | 1.808643 | -4.736757 | 9.629960 | 10 | False |
| 10 | donor_13 | 3.392449 | 5.417947 | 1.597061 | 4.111628 | 8.678758 | 11 | False |
| 11 | treated | 3.631225 | 5.591349 | 1.539797 | 4.796689 | 8.149984 | 12 | True |
| 12 | donor_6 | 2.935853 | 4.298823 | 1.464250 | 2.850820 | 8.233900 | 13 | False |
| 13 | donor_20 | 5.842398 | 8.411978 | 1.439816 | 5.493484 | 14.653821 | 14 | False |
| 14 | donor_17 | 4.721009 | 6.460381 | 1.368432 | 5.181488 | 9.619748 | 15 | False |
| 15 | donor_5 | 3.678624 | 4.950369 | 1.345712 | 2.184354 | 9.502808 | 16 | False |
| 16 | donor_3 | 6.904353 | 9.109156 | 1.319335 | 9.081197 | 9.785759 | 17 | False |
| 17 | donor_19 | 2.885092 | 3.683051 | 1.276580 | -3.615034 | 4.361489 | 18 | False |
| 18 | donor_10 | 3.852226 | 4.589747 | 1.191453 | 0.328234 | 7.374027 | 19 | False |
| 19 | donor_16 | 2.681441 | 2.057474 | 0.767302 | -0.852681 | 2.657123 | 20 | False |
| 20 | donor_9 | 3.830376 | 2.389494 | 0.623827 | -0.636729 | 3.154982 | 21 | False |
How to read columns:
- unit_id: unit used as treated in that placebo run.
- pre_rmse: pre-treatment fit error (lower is better).
- post_rmse: post-treatment fit error.
- post_pre_rmspe_ratio: post_rmse / pre_rmse (main Abadie-style statistic).
- average_post_gap: mean post-treatment gap (observed - synthetic).
- max_abs_post_gap: largest absolute post gap.
- rank_post_pre_rmspe_ratio: rank of ratio, descending (1 = largest ratio).
- is_actual_treated: True for your real treated unit.
Interpretation: You want the real treated unit (is_actual_treated=True) to have one of the largest post_pre_rmspe_ratio values (ideally rank near 1). If many placebo units have similar/larger ratios, treatment effect is less convincing. Also check pre_rmse: if pre-fit is poor for some placebos, their high ratios are less informative.
Sensitivity
| dropped_donor | average_att_reestimated | delta_vs_full_model | pre_rmse_reestimated | max_weight_after_refit | effective_n_donors_after_refit | |
|---|---|---|---|---|---|---|
| 0 | donor_1 | 4.058002 | -0.914725 | 3.698383 | 0.161013 | 8.102712 |
| 1 | donor_10 | 4.434911 | -0.537816 | 3.639324 | 0.219260 | 5.749925 |
| 2 | donor_11 | 3.689260 | -1.283467 | 3.688131 | 0.251006 | 5.031053 |
| 3 | donor_12 | 4.998016 | 0.025289 | 3.655190 | 0.223262 | 6.959361 |
| 4 | donor_13 | 4.487509 | -0.485218 | 3.631670 | 0.220332 | 5.983026 |
| 5 | donor_14 | 4.617818 | -0.354909 | 3.697599 | 0.213398 | 5.489285 |
| 6 | donor_15 | 3.880155 | -1.092572 | 3.657946 | 0.206193 | 6.328750 |
| 7 | donor_16 | 4.561821 | -0.410907 | 3.631450 | 0.222833 | 5.832072 |
| 8 | donor_17 | 4.260393 | -0.712334 | 3.636798 | 0.218268 | 6.296147 |
| 9 | donor_18 | 4.148347 | -0.824380 | 3.632912 | 0.221292 | 5.896571 |
| 10 | donor_19 | 4.623318 | -0.349409 | 3.631406 | 0.220039 | 5.951044 |
| 11 | donor_2 | 4.089068 | -0.883660 | 3.632867 | 0.218513 | 6.036224 |
| 12 | donor_20 | 5.000086 | 0.027359 | 3.660505 | 0.250500 | 5.511244 |
| 13 | donor_3 | 4.604093 | -0.368634 | 3.633820 | 0.223262 | 5.772354 |
| 14 | donor_4 | 4.864541 | -0.108186 | 3.652023 | 0.218231 | 6.019011 |
| 15 | donor_5 | 4.555381 | -0.417346 | 3.631324 | 0.222967 | 5.844479 |
| 16 | donor_6 | 4.205424 | -0.767303 | 3.649957 | 0.203193 | 7.324002 |
| 17 | donor_7 | 4.514112 | -0.458616 | 3.631452 | 0.224113 | 5.809195 |
| 18 | donor_8 | 4.256794 | -0.715934 | 3.638297 | 0.226399 | 5.976241 |
| 19 | donor_9 | 4.531504 | -0.441223 | 3.648459 | 0.219720 | 5.889569 |
- dropped_donor: donor unit removed in that refit.
- average_att_reestimated: ATT after removing that donor.
- delta_vs_full_model: reestimated_ATT - original_full_model_ATT. Near 0 estimate is stable to that donor. Large |deltal that donor is influential.
- pre_rmse_reestimated: pre-treatment fit error after refit. Higher than baseline worse pre-fit when donor is removed.
- max_weight_afterrefit: largest donor weight in refit.Very high (close to 1) dependence on one donor.
- effective_n_donors_after_refit: effective donor count (1 / sum (w^2)). Lower concentration on fewer donors.
Model
{'metrics': {'n_donors': 20, 'n_pre': 181, 'n_post': 4, 'pre_rmse_sc': None, 'pre_rmse_aug': 3.63162621137165, 'pre_rmse_augmented': 3.63162621137165, 'pre_mae_augmented': 2.865683953423894, 'max_abs_pre_gap': 12.319490234592525, 'max_abs_pre_gap_augmented': 12.319490234592525, 'mean_signed_pre_gap': -0.093703057672044, 'mean_signed_pre_gap_augmented': -0.093703057672044, 'att_sc': None, 'att_aug': 4.972727350333998, 'max_weight_sc': 0.1878646195458257, 'max_abs_weight_aug': 0.21641008257012256, 'max_abs_weight_augmented': 0.21641008257012256, 'l1_norm_weight_aug': 1.36881478737466, 'l1_norm_weights_augmented': 1.36881478737466, 'sum_weights_augmented': 1.0, 'effective_n_donors': None, 'effective_n_donors_sc': None, 'herfindahl_weights': None, 'herfindahl_weights_sc': None, 'effective_n_donors_abs_augmented': 11.74898939775322, 'herfindahl_abs_augmented': 0.0851137034978712, 'negative_weight_share': 0.13472048621056845, 'n_negative_weights': 5, 'cond_augmented_gram': 1051.7388011810417, 'n_placebos': 0, 'min_possible_p': None, 'p_value_att': 0.04108734188619114, 'ci_low_abs': 0.25034659065689, 'ci_high_abs': 9.695108110011105, 'placebo_ci_is_unbounded': False, 'missing_cell_fraction': None, 'completion_converged': None, 'completion_effective_rank': None, 'mean_gap_last_k_pre_sc': None, 'mean_gap_last_k_pre_aug': 1.8972574040111958, 'pre_tail_k_used': 3, 'slsqp_fallback_count': 2, 'slsqp_fallback_reasons': ['Positive directional derivative for linesearch'], 'suppressed_fit_warning_count': 1, 'suppressed_fit_warnings': ['SLSQP simplex optimization did not converge; falling back to projected-gradient solver. Occurrences: 2. Reason: Positive directional derivative for linesearch']}, 'warnings': ['SLSQP simplex optimization did not converge; falling back to projected-gradient solver. Occurrences: 2. Reason: Positive directional derivative for linesearch']}