generate_scm_poisson_26()
This notebook presents the generate scm poisson 26 research workflow and key analysis steps.
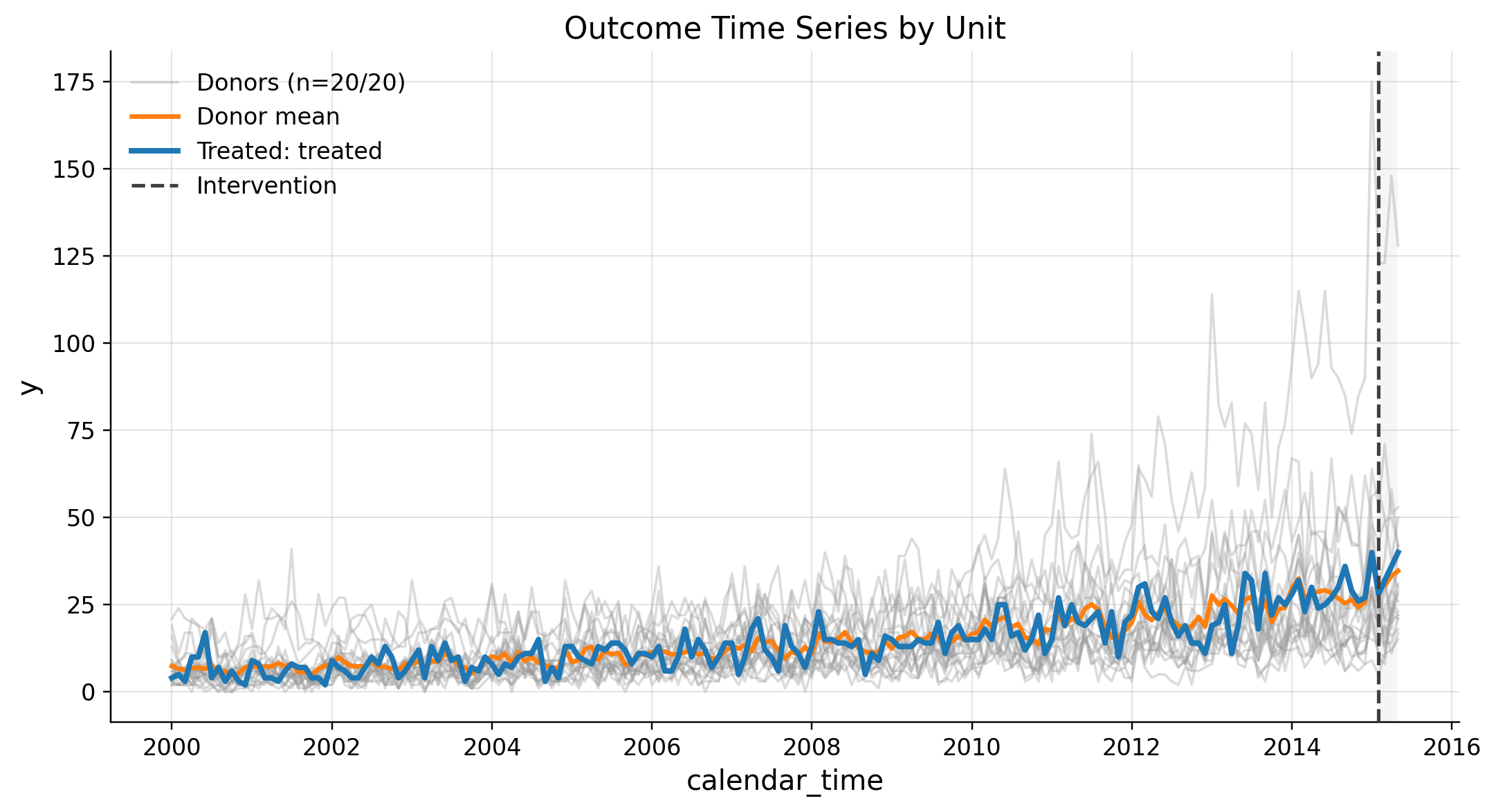
This scenario produces synthetic panel data with one treated unit and multiple donors. It uses a low-level Poisson DGP to simulate realistic discrete outcomes with time-varying exposure and latent rates.
DGP math
The Data Generating Process (DGP) for the Poisson SCM scenario follows a hierarchical log-linear model for the mean , with observations sampled from a Poisson distribution.
1.1 Donor units
For each donor unit at time :
- Exposure : where and are AR(1) processes.
- Mean :
where:
- : monthly seasonality signal
- : common factor (macro log index, AR(1))
- : latent factors (AR(1))
- : donor-specific AR(1) noise
- Outcome :
1.2 Treated unit
The counterfactual mean is a weighted combination of donors, potentially with a pre-fit mismatch: where .
The treated mean is: where follows a post-treatment ramp-in path: and for all pre-treatment and intervention-anchor periods.
The outcomes and are coupled via a thinning/superposition property to maintain exact Poisson marginals while ensuring the realized effect is driven by the multiplier in expectation:
- If :
- If :
This ensures that and both marginals remain Poisson with means and .
2. Oracle Treatment Effects (ATT)
The Average Treatment Effect on the Treated (ATT) is the average impact of the intervention across all post-treatment periods. In this synthetic scenario, we can calculate it in two ways:
-
Realized ATT: Based on observed vs. counterfactual outcomes. In the data, this is the mean of
tau_realized_truefor the treated unit in post-periods. -
Mean ATT: Based on the underlying population means (the "signal"). In the data, this is the mean of
tau_mean_truefor the treated unit in post-periods.
Ground-truth ATTE is 2.750000
| unit_id | calendar_time | treated_time | y | y_cf | tau_realized_true | mu_cf | mu_treated | tau_mean_true | |
|---|---|---|---|---|---|---|---|---|---|
| 0 | donor_1 | 2000-01 | 0 | 4.0 | 4.0 | 0.0 | 6.150671 | 6.150671 | 0.0 |
| 1 | donor_1 | 2000-02 | 0 | 9.0 | 9.0 | 0.0 | 5.599403 | 5.599403 | 0.0 |
| 2 | donor_1 | 2000-03 | 0 | 4.0 | 4.0 | 0.0 | 4.978119 | 4.978119 | 0.0 |
| 3 | donor_1 | 2000-04 | 0 | 6.0 | 6.0 | 0.0 | 5.777247 | 5.777247 | 0.0 |
| 4 | donor_1 | 2000-05 | 0 | 8.0 | 8.0 | 0.0 | 5.685711 | 5.685711 | 0.0 |
EDA
PanelDataSCM(df=(3885, 4), y='y', unit_col='unit_id', time_col='calendar_time', treated_time='treated_time', time_freq='M', treated_unit='treated', treatment_start=Period('2015-02', 'M'), last_post_period=Period('2015-05', 'M'), n_pre_periods=181, n_post_periods=4, donor_units=['donor_1', 'donor_10', 'donor_11', 'donor_12', 'donor_13', 'donor_14', 'donor_15', 'donor_16', 'donor_17', 'donor_18', 'donor_19', 'donor_2', 'donor_20', 'donor_3', 'donor_4', 'donor_5', 'donor_6', 'donor_7', 'donor_8', 'donor_9'])
| donor | pre_mean | pre_std | pre_slope | corr_with_treated_pre | rmse_to_treated_pre | rmse_to_treated_pre_standardized | mean_diff_pre | slope_diff_pre | max_abs_gap_pre | is_never_treated | n_missing_pre | corr_rank | std_rmse_rank | slope_rank | composite_similarity_score | rank_by_similarity | notes | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | donor_4 | 9.762431 | 7.179154 | 0.115553 | 0.775096 | 6.670625 | 0.842433 | -4.287293 | -0.007929 | 20.0 | True | 0 | 4 | 5 | 1 | 0.883333 | 1 | ok |
| 1 | donor_10 | 13.563536 | 7.602892 | 0.114674 | 0.755004 | 5.462074 | 0.689805 | -0.486188 | -0.008807 | 17.0 | True | 0 | 8 | 1 | 2 | 0.866667 | 2 | ok |
| 2 | donor_18 | 16.220994 | 9.298357 | 0.140931 | 0.774548 | 6.310125 | 0.796906 | 2.171271 | 0.017449 | 20.0 | True | 0 | 5 | 3 | 3 | 0.866667 | 3 | ok |
| 3 | donor_9 | 13.701657 | 6.733231 | 0.100152 | 0.731460 | 5.491832 | 0.693564 | -0.348066 | -0.023330 | 19.0 | True | 0 | 10 | 2 | 4 | 0.783333 | 4 | ok |
| 4 | donor_1 | 10.646409 | 5.490006 | 0.079764 | 0.740206 | 6.329794 | 0.799390 | -3.403315 | -0.043717 | 20.0 | True | 0 | 9 | 4 | 6 | 0.733333 | 5 | ok |
| 5 | donor_17 | 17.458564 | 10.886897 | 0.178779 | 0.770667 | 7.744183 | 0.978013 | 3.408840 | 0.055297 | 27.0 | True | 0 | 6 | 9 | 9 | 0.650000 | 6 | ok |
| 6 | donor_19 | 10.453039 | 5.153367 | 0.073788 | 0.691311 | 6.765610 | 0.854429 | -3.596685 | -0.049693 | 23.0 | True | 0 | 12 | 6 | 7 | 0.633333 | 7 | ok |
| 7 | donor_13 | 8.718232 | 5.794182 | 0.080950 | 0.681039 | 7.886978 | 0.996047 | -5.331492 | -0.042531 | 30.0 | True | 0 | 13 | 10 | 5 | 0.583333 | 8 | ok |
| 8 | donor_8 | 9.198895 | 5.152288 | 0.070184 | 0.669963 | 7.623024 | 0.962712 | -4.850829 | -0.053298 | 23.0 | True | 0 | 14 | 8 | 8 | 0.550000 | 9 | ok |
| 9 | donor_5 | 10.950276 | 5.267723 | 0.065347 | 0.609616 | 7.014192 | 0.885823 | -3.099448 | -0.058134 | 22.0 | True | 0 | 16 | 7 | 10 | 0.500000 | 10 | ok |
| 10 | donor_7 | 7.204420 | 4.610483 | 0.064872 | 0.696173 | 8.943345 | 1.129455 | -6.845304 | -0.058610 | 22.0 | True | 0 | 11 | 12 | 11 | 0.483333 | 11 | high_std_rmse |
| 11 | donor_11 | 21.038674 | 12.944576 | 0.224528 | 0.828828 | 10.449828 | 1.319709 | 6.988950 | 0.101046 | 35.0 | True | 0 | 2 | 16 | 17 | 0.466667 | 12 | high_std_rmse |
| 12 | donor_14 | 15.685083 | 15.073494 | 0.250056 | 0.832498 | 9.687949 | 1.223491 | 1.635359 | 0.126574 | 34.0 | True | 0 | 1 | 15 | 19 | 0.466667 | 13 | high_std_rmse |
| 13 | donor_6 | 8.690608 | 4.149602 | 0.049649 | 0.564090 | 8.459850 | 1.068395 | -5.359116 | -0.073833 | 27.0 | True | 0 | 17 | 11 | 13 | 0.366667 | 14 | high_std_rmse |
| 14 | donor_12 | 7.569061 | 3.764025 | 0.046441 | 0.618555 | 9.055080 | 1.143566 | -6.480663 | -0.077040 | 31.0 | True | 0 | 15 | 13 | 14 | 0.350000 | 15 | high_std_rmse |
| 15 | donor_15 | 27.751381 | 13.661215 | 0.212821 | 0.764745 | 16.480694 | 2.081347 | 13.701657 | 0.089339 | 53.0 | True | 0 | 7 | 19 | 16 | 0.350000 | 16 | high_std_rmse |
| 16 | donor_2 | 31.861878 | 29.367320 | 0.482832 | 0.825508 | 29.299935 | 3.700290 | 17.812155 | 0.359351 | 135.0 | True | 0 | 3 | 20 | 20 | 0.333333 | 17 | high_std_rmse |
| 17 | donor_20 | 19.972376 | 6.508330 | 0.056204 | 0.480955 | 9.516488 | 1.201837 | 5.922652 | -0.067278 | 23.0 | True | 0 | 19 | 14 | 12 | 0.300000 | 18 | high_std_rmse |
| 18 | donor_16 | 5.662983 | 3.652322 | 0.040390 | 0.561087 | 10.673455 | 1.347951 | -8.386740 | -0.083091 | 31.0 | True | 0 | 18 | 17 | 15 | 0.216667 | 19 | high_std_rmse |
| 19 | donor_3 | 19.900552 | 5.562408 | -0.000138 | 0.105856 | 10.887927 | 1.375037 | 5.850829 | -0.123619 | 33.0 | True | 0 | 20 | 18 | 18 | 0.116667 | 20 | high_std_rmse |