generate_scm_gamma_26()
This notebook presents the generate scm gamma 26 research workflow and key analysis steps.
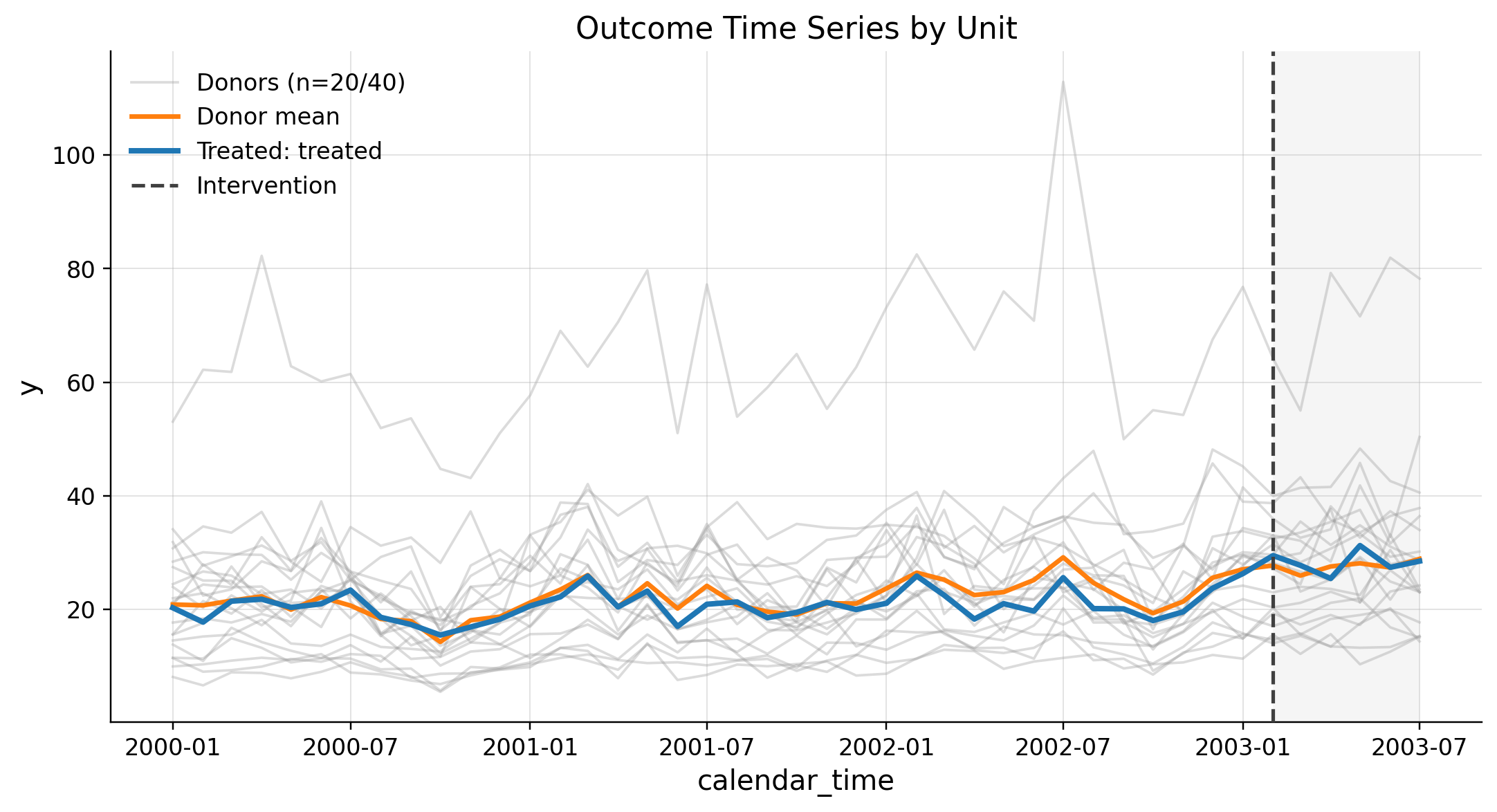
This scenario produces synthetic panel data with one treated unit and multiple donors. It uses a low-level Gamma DGP to simulate realistic outcomes with time-varying exposure and latent rates.
DGP math
The Data Generating Process (DGP) for the Gamma SCM scenario follows a hierarchical log-linear model for the mean , with observations sampled from a Gamma distribution.
1.1 Donor units
For each donor unit at time :
- Exposure : where and are AR(1) processes.
- Mean :
where:
- : monthly seasonality signal
- : common factor (macro log index, AR(1))
- : latent factors (AR(1))
- : donor-specific AR(1) noise
- Outcome :
where is the
gamma_shapeparameter.
1.2 Treated unit
The counterfactual mean is a weighted combination of donors, potentially with a pre-fit mismatch: where .
Treatment is active only in post periods. For post step :
where is treatment_effect_rate and is treatment_effect_slope.
Equivalently, in calendar time:
The outcomes are: So for each , remains Gamma-distributed with mean , and is exactly the realized treatment effect.
2. Oracle Treatment Effects (ATT)
The Average Treatment Effect on the Treated (ATT) is the average impact of the intervention across all post-treatment periods. In this synthetic scenario, we can calculate it in two ways:
-
Realized ATT: Based on observed vs. counterfactual outcomes. In the data, this is the mean of
tau_realized_truefor the treated unit in post-periods. -
Mean ATT: Based on the underlying population means (the "signal"). In the data, this is the mean of
tau_mean_truefor the treated unit in post-periods.
Ground-truth ATTE is 1.919269
| unit_id | calendar_time | treated_time | y | y_cf | tau_realized_true | mu_cf | mu_treated | tau_mean_true | |
|---|---|---|---|---|---|---|---|---|---|
| 0 | donor_1 | 2000-01 | 0 | 9.971611 | 9.971611 | 0.0 | 10.059081 | 10.059081 | 0.0 |
| 1 | donor_1 | 2000-02 | 0 | 10.344379 | 10.344379 | 0.0 | 10.097467 | 10.097467 | 0.0 |
| 2 | donor_1 | 2000-03 | 0 | 10.998498 | 10.998498 | 0.0 | 11.354098 | 11.354098 | 0.0 |
| 3 | donor_1 | 2000-04 | 0 | 11.508717 | 11.508717 | 0.0 | 11.277716 | 11.277716 | 0.0 |
| 4 | donor_1 | 2000-05 | 0 | 11.125281 | 11.125281 | 0.0 | 10.098791 | 10.098791 | 0.0 |
EDA
PanelDataSCM(df=(1763, 4), y='y', unit_col='unit_id', time_col='calendar_time', treated_time='treated_time', time_freq='M', treated_unit='treated', treatment_start=Period('2003-02', 'M'), last_post_period=Period('2003-07', 'M'), n_pre_periods=37, n_post_periods=6, donor_units=['donor_1', 'donor_10', 'donor_11', 'donor_12', 'donor_13', 'donor_14', 'donor_15', 'donor_16', 'donor_17', 'donor_18', 'donor_19', 'donor_2', 'donor_20', 'donor_21', 'donor_22', 'donor_23', 'donor_24', 'donor_25', 'donor_26', 'donor_27', 'donor_28', 'donor_29', 'donor_3', 'donor_30', 'donor_31', 'donor_32', 'donor_33', 'donor_34', 'donor_35', 'donor_36', 'donor_37', 'donor_38', 'donor_39', 'donor_4', 'donor_40', 'donor_5', 'donor_6', 'donor_7', 'donor_8', 'donor_9'])
| donor | pre_mean | pre_std | pre_slope | corr_with_treated_pre | rmse_to_treated_pre | rmse_to_treated_pre_standardized | mean_diff_pre | slope_diff_pre | max_abs_gap_pre | is_never_treated | n_missing_pre | corr_rank | std_rmse_rank | slope_rank | composite_similarity_score | rank_by_similarity | notes | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | donor_20 | 20.756877 | 4.235256 | 0.065672 | 0.699298 | 3.050574 | 1.177242 | 0.055800 | 0.001045 | 7.515230 | True | 0 | 6 | 2 | 1 | 0.950000 | 1 | high_std_rmse |
| 1 | donor_23 | 18.859982 | 3.459160 | 0.109325 | 0.792879 | 2.802849 | 1.081642 | -1.841096 | 0.044698 | 6.193337 | True | 0 | 1 | 1 | 19 | 0.850000 | 2 | high_std_rmse |
| 2 | donor_17 | 27.374399 | 5.004251 | 0.062159 | 0.694945 | 7.633294 | 2.945751 | 6.673322 | -0.002467 | 15.022205 | True | 0 | 7 | 23 | 3 | 0.750000 | 3 | high_std_rmse |
| 3 | donor_35 | 12.890196 | 2.478484 | 0.054896 | 0.774341 | 7.995072 | 3.085364 | -7.810882 | -0.009730 | 11.244939 | True | 0 | 2 | 25 | 7 | 0.741667 | 4 | high_std_rmse |
| 4 | donor_27 | 12.138052 | 2.561773 | 0.065732 | 0.710107 | 8.784934 | 3.390178 | -8.563025 | 0.001106 | 12.226100 | True | 0 | 4 | 29 | 2 | 0.733333 | 5 | high_std_rmse |
| 5 | donor_39 | 15.933235 | 3.493479 | 0.101601 | 0.756099 | 5.287926 | 2.040654 | -4.767843 | 0.036974 | 9.398086 | True | 0 | 3 | 16 | 16 | 0.733333 | 6 | high_std_rmse |
| 6 | donor_2 | 22.023910 | 3.752940 | 0.025593 | 0.599773 | 3.299044 | 1.273128 | 1.322833 | -0.039033 | 8.913091 | True | 0 | 19 | 4 | 17 | 0.691667 | 7 | high_std_rmse |
| 7 | donor_34 | 23.225266 | 4.258988 | 0.116799 | 0.685302 | 4.012347 | 1.548398 | 2.524189 | 0.052173 | 8.782578 | True | 0 | 8 | 11 | 21 | 0.691667 | 8 | high_std_rmse |
| 8 | donor_26 | 15.531548 | 2.367240 | 0.058569 | 0.584382 | 5.645636 | 2.178697 | -5.169529 | -0.006057 | 10.132082 | True | 0 | 21 | 18 | 5 | 0.658333 | 9 | high_std_rmse |
| 9 | donor_9 | 20.198346 | 3.894696 | 0.185173 | 0.609819 | 3.134843 | 1.209762 | -0.502731 | 0.120546 | 10.776096 | True | 0 | 15 | 3 | 31 | 0.616667 | 10 | high_std_rmse |
| 10 | donor_31 | 15.974505 | 3.809154 | 0.046519 | 0.573221 | 5.678805 | 2.191497 | -4.726572 | -0.018107 | 12.461670 | True | 0 | 22 | 19 | 9 | 0.608333 | 11 | high_std_rmse |
| 11 | donor_3 | 22.984470 | 4.678813 | 0.162331 | 0.639562 | 4.279210 | 1.651382 | 2.283392 | 0.097704 | 12.714594 | True | 0 | 12 | 13 | 26 | 0.600000 | 12 | high_std_rmse |
| 12 | donor_7 | 19.888000 | 3.565032 | 0.048846 | 0.434870 | 3.471405 | 1.339644 | -0.813077 | -0.015780 | 8.749577 | True | 0 | 37 | 6 | 8 | 0.600000 | 13 | high_std_rmse |
| 13 | donor_33 | 22.467401 | 3.890376 | 0.061634 | 0.437267 | 4.019136 | 1.551018 | 1.766323 | -0.002993 | 9.071825 | True | 0 | 36 | 12 | 4 | 0.591667 | 14 | high_std_rmse |
| 14 | donor_6 | 10.817057 | 1.619471 | 0.030223 | 0.705904 | 10.055180 | 3.880377 | -9.884020 | -0.034404 | 15.388984 | True | 0 | 5 | 33 | 14 | 0.591667 | 15 | high_std_rmse |
| 15 | donor_21 | 21.001540 | 4.037106 | -0.018281 | 0.542038 | 3.429640 | 1.323526 | 0.300463 | -0.082908 | 10.637405 | True | 0 | 27 | 5 | 25 | 0.550000 | 16 | high_std_rmse |
| 16 | donor_13 | 28.400664 | 5.292688 | 0.037227 | 0.605290 | 8.798181 | 3.395290 | 7.699587 | -0.027399 | 17.004455 | True | 0 | 18 | 30 | 12 | 0.525000 | 17 | high_std_rmse |
| 17 | donor_15 | 21.647789 | 4.210829 | 0.182164 | 0.554289 | 3.639493 | 1.404510 | 0.946712 | 0.117537 | 13.105415 | True | 0 | 24 | 7 | 29 | 0.525000 | 18 | high_std_rmse |
| 18 | donor_12 | 9.964996 | 1.894391 | 0.090093 | 0.607253 | 10.936409 | 4.220450 | -10.736082 | 0.025466 | 14.965497 | True | 0 | 17 | 34 | 10 | 0.516667 | 19 | high_std_rmse |
| 19 | donor_8 | 8.772232 | 2.098724 | 0.104214 | 0.647465 | 12.098529 | 4.668922 | -11.928845 | 0.039588 | 17.063234 | True | 0 | 10 | 35 | 18 | 0.500000 | 20 | high_std_rmse |
| 20 | donor_30 | 23.684935 | 4.356977 | 0.116176 | 0.497646 | 4.833674 | 1.865355 | 2.983858 | 0.051549 | 10.057305 | True | 0 | 30 | 15 | 20 | 0.483333 | 21 | high_std_rmse |
| 21 | donor_4 | 21.801573 | 4.207243 | 0.179567 | 0.524523 | 3.766945 | 1.453695 | 1.100496 | 0.114941 | 8.288093 | True | 0 | 28 | 9 | 28 | 0.483333 | 22 | high_std_rmse |
| 22 | donor_10 | 22.140468 | 4.508805 | 0.224646 | 0.558779 | 4.007346 | 1.546468 | 1.439391 | 0.160019 | 9.740145 | True | 0 | 23 | 10 | 33 | 0.475000 | 23 | high_std_rmse |
| 23 | donor_5 | 25.412361 | 4.374273 | 0.057271 | 0.323830 | 6.379970 | 2.462083 | 4.711284 | -0.007355 | 14.186181 | True | 0 | 40 | 21 | 6 | 0.466667 | 24 | high_std_rmse |
| 24 | donor_18 | 15.375511 | 2.894278 | 0.142564 | 0.551353 | 5.931533 | 2.289027 | -5.325566 | 0.077938 | 11.432539 | True | 0 | 25 | 20 | 23 | 0.458333 | 25 | high_std_rmse |
| 25 | donor_25 | 25.638145 | 4.719678 | 0.100007 | 0.491059 | 6.430663 | 2.481646 | 4.937068 | 0.035381 | 13.366836 | True | 0 | 33 | 22 | 15 | 0.441667 | 26 | high_std_rmse |
| 26 | donor_36 | 17.556815 | 3.510483 | 0.132070 | 0.490282 | 4.472671 | 1.726041 | -3.144262 | 0.067444 | 10.183903 | True | 0 | 34 | 14 | 22 | 0.441667 | 27 | high_std_rmse |
| 27 | donor_1 | 11.442375 | 2.525462 | 0.146053 | 0.608511 | 9.531624 | 3.678332 | -9.258702 | 0.081427 | 14.575727 | True | 0 | 16 | 31 | 24 | 0.433333 | 28 | high_std_rmse |
| 28 | donor_40 | 11.363776 | 1.958071 | 0.090476 | 0.506775 | 9.622439 | 3.713378 | -9.337301 | 0.025850 | 14.358223 | True | 0 | 29 | 32 | 11 | 0.425000 | 29 | high_std_rmse |
| 29 | donor_32 | 18.875009 | 3.180241 | -0.049527 | 0.404701 | 3.673285 | 1.417551 | -1.826069 | -0.114153 | 9.321772 | True | 0 | 38 | 8 | 27 | 0.416667 | 30 | high_std_rmse |
| 30 | donor_28 | 27.821562 | 5.599506 | 0.336526 | 0.645032 | 8.369695 | 3.229934 | 7.120484 | 0.271899 | 16.331011 | True | 0 | 11 | 26 | 37 | 0.408333 | 31 | high_std_rmse |
| 31 | donor_16 | 26.230599 | 5.760959 | 0.095907 | 0.370933 | 7.707416 | 2.974355 | 5.529522 | 0.031281 | 18.330394 | True | 0 | 39 | 24 | 13 | 0.391667 | 32 | high_std_rmse |
| 32 | donor_11 | 31.898384 | 5.748397 | 0.278022 | 0.627211 | 12.101775 | 4.670174 | 11.197307 | 0.213395 | 24.414887 | True | 0 | 13 | 36 | 35 | 0.325000 | 33 | high_std_rmse |
| 33 | donor_38 | 23.341229 | 5.607884 | 0.327023 | 0.473125 | 5.602050 | 2.161877 | 2.640152 | 0.262396 | 13.541965 | True | 0 | 35 | 17 | 36 | 0.291667 | 34 | high_std_rmse |
| 34 | donor_19 | 64.642074 | 13.285880 | 0.416100 | 0.666003 | 45.477279 | 17.550056 | 43.940997 | 0.351474 | 87.274045 | True | 0 | 9 | 40 | 40 | 0.283333 | 35 | high_std_rmse |
| 35 | donor_29 | 32.069172 | 6.680389 | 0.392642 | 0.623576 | 12.608994 | 4.865914 | 11.368095 | 0.328015 | 20.594296 | True | 0 | 14 | 37 | 38 | 0.283333 | 36 | high_std_rmse |
| 36 | donor_24 | 12.604643 | 3.169244 | 0.199055 | 0.493436 | 8.614321 | 3.324337 | -8.096434 | 0.134428 | 14.907594 | True | 0 | 31 | 28 | 32 | 0.266667 | 37 | high_std_rmse |
| 37 | donor_14 | 27.342267 | 6.286205 | 0.393100 | 0.547435 | 8.514798 | 3.285931 | 6.641190 | 0.328473 | 15.190467 | True | 0 | 26 | 27 | 39 | 0.258333 | 38 | high_std_rmse |
| 38 | donor_37 | 34.405731 | 7.663463 | 0.231876 | 0.590564 | 15.159349 | 5.850117 | 13.704654 | 0.167249 | 33.571787 | True | 0 | 20 | 39 | 34 | 0.250000 | 39 | high_std_rmse |
| 39 | donor_22 | 34.353351 | 5.235590 | 0.183591 | 0.492258 | 14.392843 | 5.554316 | 13.652274 | 0.118964 | 27.762663 | True | 0 | 32 | 38 | 30 | 0.191667 | 40 | high_std_rmse |